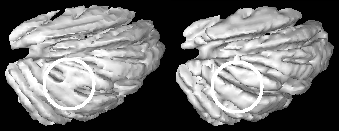
Automatic Surface Extraction by Discrete, Topologically Controlled,
Region Growing
Josh Stern

 ,
Kirt Schaper
,
Kirt Schaper ,
Kelly Rehm
,
Kelly Rehm
 ,
Monica Hurdal
,
Monica Hurdal ,
De Witt Sumners
,
De Witt Sumners ,
,David Rottenberg

 Department of Neurology, University of Minnesota
Department of Neurology, University of Minnesota PET Imaging Center, Minneapolis VA Medical Center
PET Imaging Center, Minneapolis VA Medical Center Department of Radiology, University of Minnesota
Department of Radiology, University of Minnesota Department of Mathematics, Florida State University
Department of Mathematics, Florida State University