Motivation:
There are various kinds of complex systems. Describing and reassembling complex systems in mathematical models is a challenging problem. Networks are a natural representation of complex systems. Therefore, network analysis is pretty important to solve the challenging problem.
A number of recent studies have focused on the statistical properties of networked systems. One common property of many networks is called the community structure. The procedure to find the community structures is "community detection".
Community Structure:
Community structure is one common property of many networks, it is the division of network nodes into groups within which the network connections are dense, but between which are sparser. These groups are called communities, or modules.
A figurative sketch of a network with such a community structure is shown as follows:
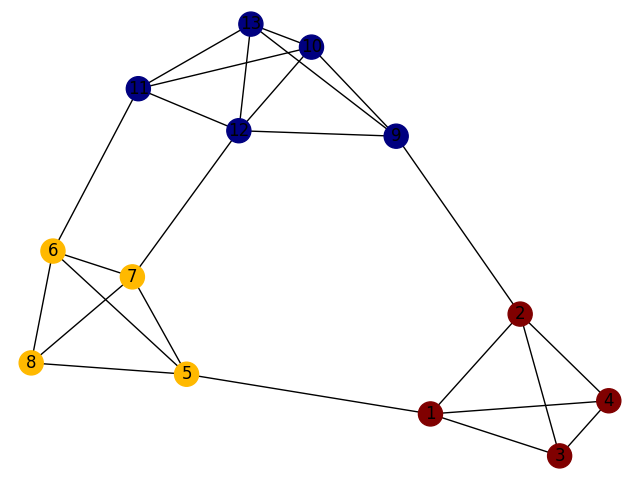
As we know, clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster) are more similar (in some sense) to each other than to those in other groups (clusters). Community detection is one specific kind of clustering(cluster analysis) and community detection is only based on networks(denoted by graphs).
Methods for Community Detection:
Minimum-cut method:
A minimum cut of a graph is a cut (a partition of the vertices of a graph into two disjoint subsets that are joined by at least one edge) that is minimal in some sense.
Edge-Betweenness method:
The Girvan-Newman algorithm detects communities by progressively removing edges from the original network based on the edge-betweenness, where the "edge betweenness" of an edge as the number of shortest paths between pairs of nodes that run along it. The algorithm's steps are as follows:
- The betweenness of all existing edges in the network is calculated first.
- The edge with the highest betweenness is removed.
- The betweenness of all edges affected by the removal is recalculated.
- Steps 2 and 3 are repeated until no edges remain.
Modularity-based method:
GN Modularity: $$Q=(\text{fraction of edges within communities)} - \text{(expected fraction of such edges}).$$
The optimization problem is $ max_{\sigma} Q(\sigma)$.
Reichardt and Bornholdt generalized the modularity function:
The generalized quality function is as follows: $$H(\sigma)=-\sum_{i,j}{[a_{ij}A_{ij}-b_{ij}(1-A_{ij})]\delta(\sigma_i,\sigma_j)+[c_{ij}A_{ij}-d_{ij}(1-A_{ij})](1-\delta(\sigma_i,\sigma_j))},$$ where $i$ and $j$ are two nodes, $A_{ij}$ is an element of the adjacency matrix, $\delta$ is the Kronecker delta symbol and $\sigma_i$ is the label of the community to which node $i$ is assigned.
The optimization problem is $ min_{\sigma} H(\sigma) $.
Statistical inference:
- The machinery of probabilistic mixture models
- The expectation-maximization algorithm
Benchmarks:
To test whether the algorithms and methods work efficiently, we need to construct some good test networks and measures.
- Two common benchamark networks;
- Newman's benchmark network: $q=4$ communities of $n_c=32$ nodes each, with $\mu$ varying from 0 to 1
- To make sure the benchmark network is close to the realistic networks, the LFR benchmark is introduced by Lancichinetti et al.
- Various measures
- Normalized mutual information (NMI)
- Variation of information (VI)
- Rand index
- Jaccard index